'data.frame': 644 obs. of 9 variables:
$ site : Ord.factor w/ 23 levels "R-1"<"R-2"<"R-3"<..: 13 14 15 1 2 3 4 5 6 7 ...
$ mined : Factor w/ 2 levels "yes","no": 1 1 1 2 2 2 2 2 2 2 ...
$ cover : num -1.442 0.298 0.398 -0.448 0.597 ...
$ sample: int 1 1 1 1 1 1 1 1 1 1 ...
$ DOP : num -0.596 -0.596 -1.191 0 0.596 ...
$ Wtemp : num -1.2294 0.0848 1.0142 -3.0234 -0.1443 ...
$ DOY : num -1.497 -1.497 -1.294 -2.712 -0.687 ...
$ spp : Factor w/ 7 levels "GP","PR","DM",..: 1 1 1 1 1 1 1 1 1 1 ...
$ count : int 0 0 0 2 2 1 1 2 4 1 ...📨 ZIP Code 📬
A hands-on R session on Zero-Inflated Poisson and other models for rare primate behaviors
João d’Oliveira Coelho
2025-04-02
Why count data is different
- Count data is discrete, non-negative, often skewed
- Examples in primatology:
- Vocalization frequencies in vervets (few 0s)
- Tool use occurrences in chimpanzees (some 0s)
- Bipedalism bouts in chacma baboons (many 0s)
- Why not use linear regression?
- Violations of assumptions: non-normality, non-constant variance
The Poisson distribution
- Poisson models count data where mean = variance
- Probability mass function: \[ P(Y = k) = \frac{\lambda^k e^{-\lambda}}{k!} \]
- Example I wanted: capuchin predation events
- Example you get: Salamanders {glmmTMB} dataset
Price SJ, Muncy BL, Bonner SJ, Drayer AN, Barton CD (2016) Effects of mountaintop removal mining and valley filling on the occupancy and abundance of stream salamanders. Journal of Applied Ecology 53 459–468. doi:10.1111/1365-2664.12585
The Poisson distribution (simulation)
lambda <- 2 # Mean and variance
pois_sample <- rpois(1000, lambda) # Random generation of Poisson distribution
ggplot(data.frame(pois_sample), aes(x = as.factor(pois_sample))) +
geom_histogram(stat = "count") + theme_minimal()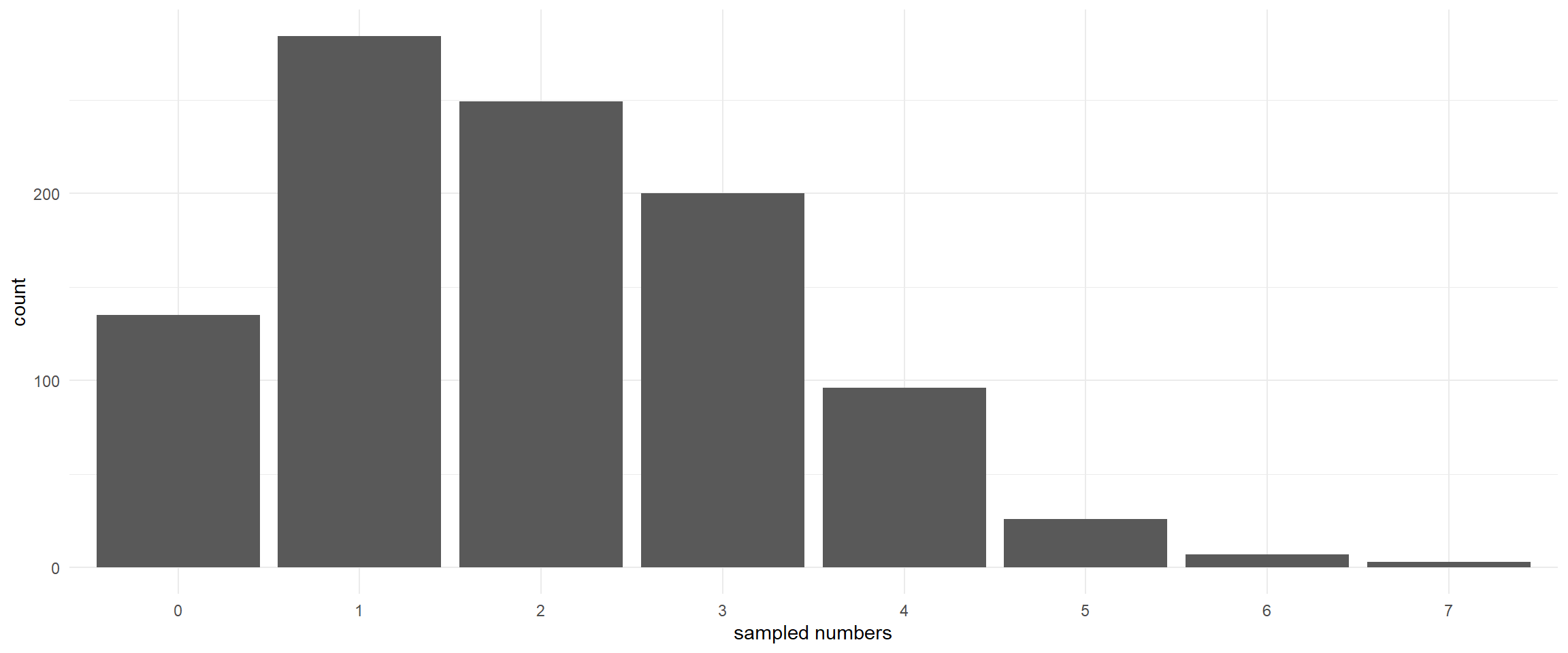
Salamander dataset
# Install if necessary
install.packages(c("glmmTMB", "ggplot2", "DHARMa"))
# Load libraries
library(glmmTMB) # For modeling
library(ggplot2) # For visualizations
library(DHARMa) # For residual diagnostics
data(Salamanders)
str(Salamanders)Salamander dataset (distribution)
[1] 0.6009317ggplot(Salamanders, aes(x = count)) +
geom_histogram(binwidth = 1) +
theme_minimal() + labs(x = "Specimen count", y = "Frequency")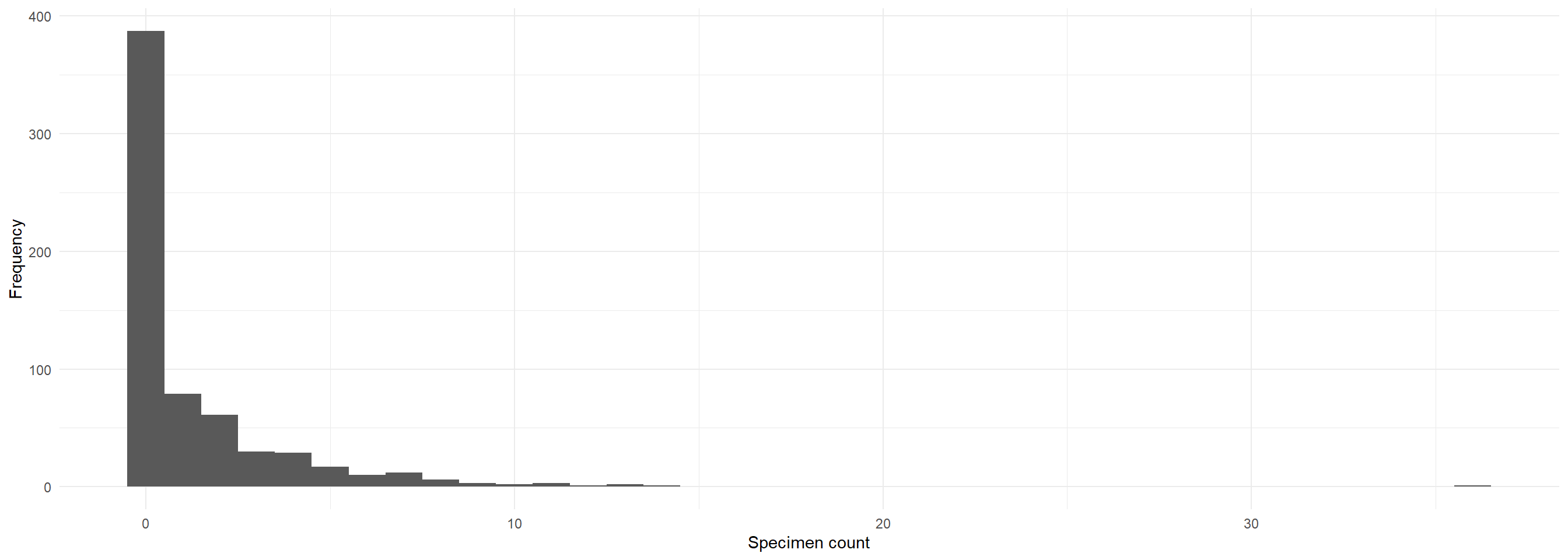
Poisson model
m1_pr <- glmmTMB(count ~ spp + mined + (1|site), data = Salamanders, family = poisson)
summary(m1_pr) Family: poisson ( log )
Formula: count ~ spp + mined + (1 | site)
Data: Salamanders
AIC BIC logLik deviance df.resid
1962.8 2003.0 -972.4 1944.8 635
Random effects:
Conditional model:
Groups Name Variance Std.Dev.
site (Intercept) 0.3316 0.5759
Number of obs: 644, groups: site, 23
Conditional model:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -1.62490 0.24048 -6.757 1.41e-11 ***
sppPR -1.38626 0.21516 -6.443 1.17e-10 ***
sppDM 0.23054 0.12889 1.789 0.0737 .
sppEC-A -0.77010 0.17105 -4.502 6.73e-06 ***
sppEC-L 0.62118 0.11931 5.207 1.92e-07 ***
sppDES-L 0.67918 0.11813 5.750 8.95e-09 ***
sppDF 0.08005 0.13344 0.600 0.5486
minedno 2.26444 0.28026 8.080 6.48e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Diagnosis of the Poisson model
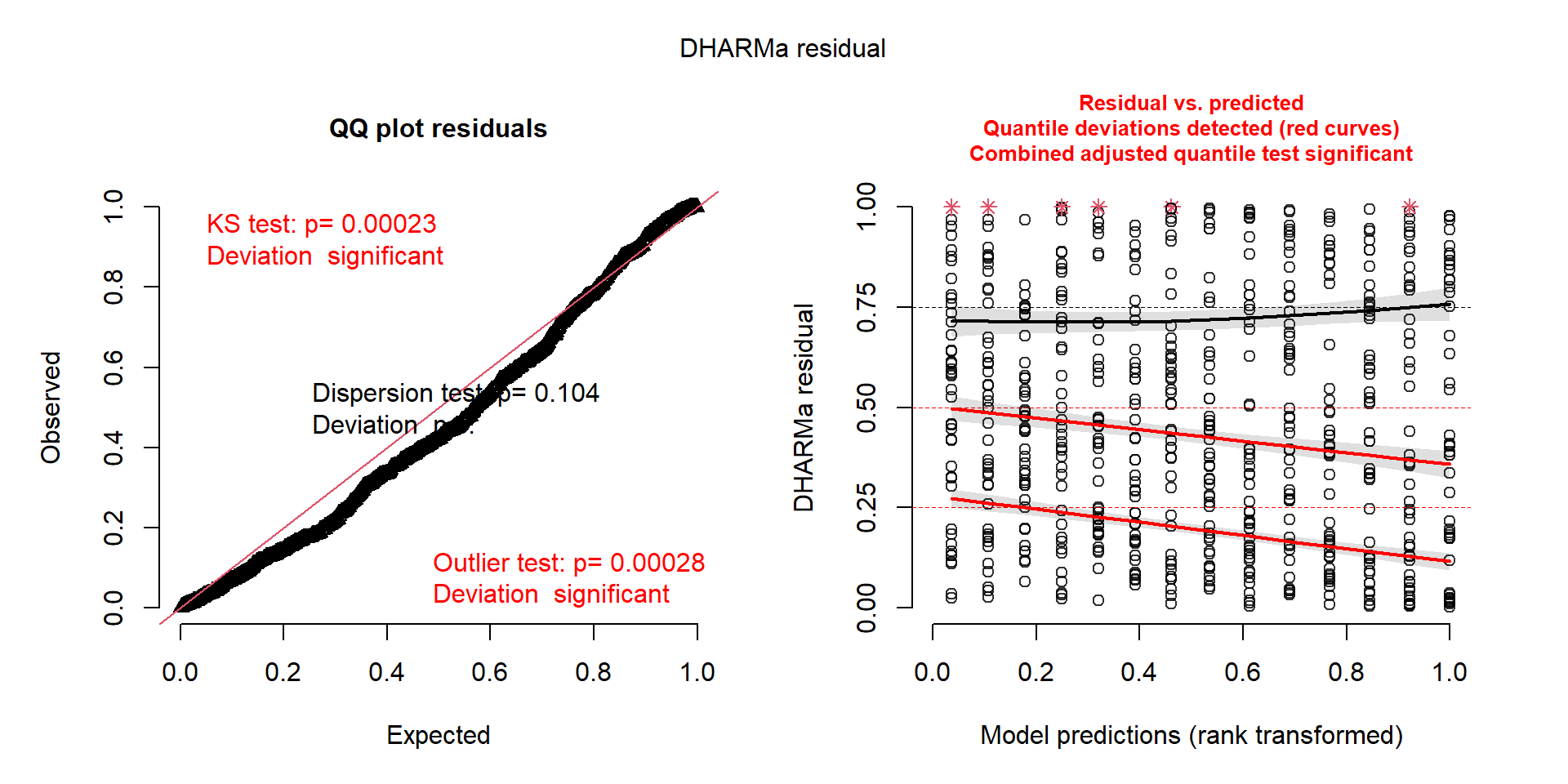
How to handle overdispersion?
- Overdispersion: Variance > Mean
- Poisson assumption often fails → underestimates SE, misleading p-values
- Negative Binomial adds a dispersion parameter to correct for this
Negative Binomial model
m2_nb <- glmmTMB(count ~ spp + mined + (1|site), data = Salamanders, family = nbinom2)
summary(m2_nb) Family: nbinom2 ( log )
Formula: count ~ spp + mined + (1 | site)
Data: Salamanders
AIC BIC logLik deviance df.resid
1672.4 1717.1 -826.2 1652.4 634
Random effects:
Conditional model:
Groups Name Variance Std.Dev.
site (Intercept) 0.2945 0.5426
Number of obs: 644, groups: site, 23
Dispersion parameter for nbinom2 family (): 0.942
Conditional model:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -1.6832 0.2742 -6.140 8.28e-10 ***
sppPR -1.3197 0.2875 -4.591 4.42e-06 ***
sppDM 0.3686 0.2235 1.649 0.099056 .
sppEC-A -0.7098 0.2530 -2.806 0.005017 **
sppEC-L 0.5714 0.2191 2.608 0.009105 **
sppDES-L 0.7929 0.2166 3.660 0.000252 ***
sppDF 0.3120 0.2329 1.340 0.180337
minedno 2.2633 0.2838 7.975 1.53e-15 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Diagnosis of the Negative Binomial
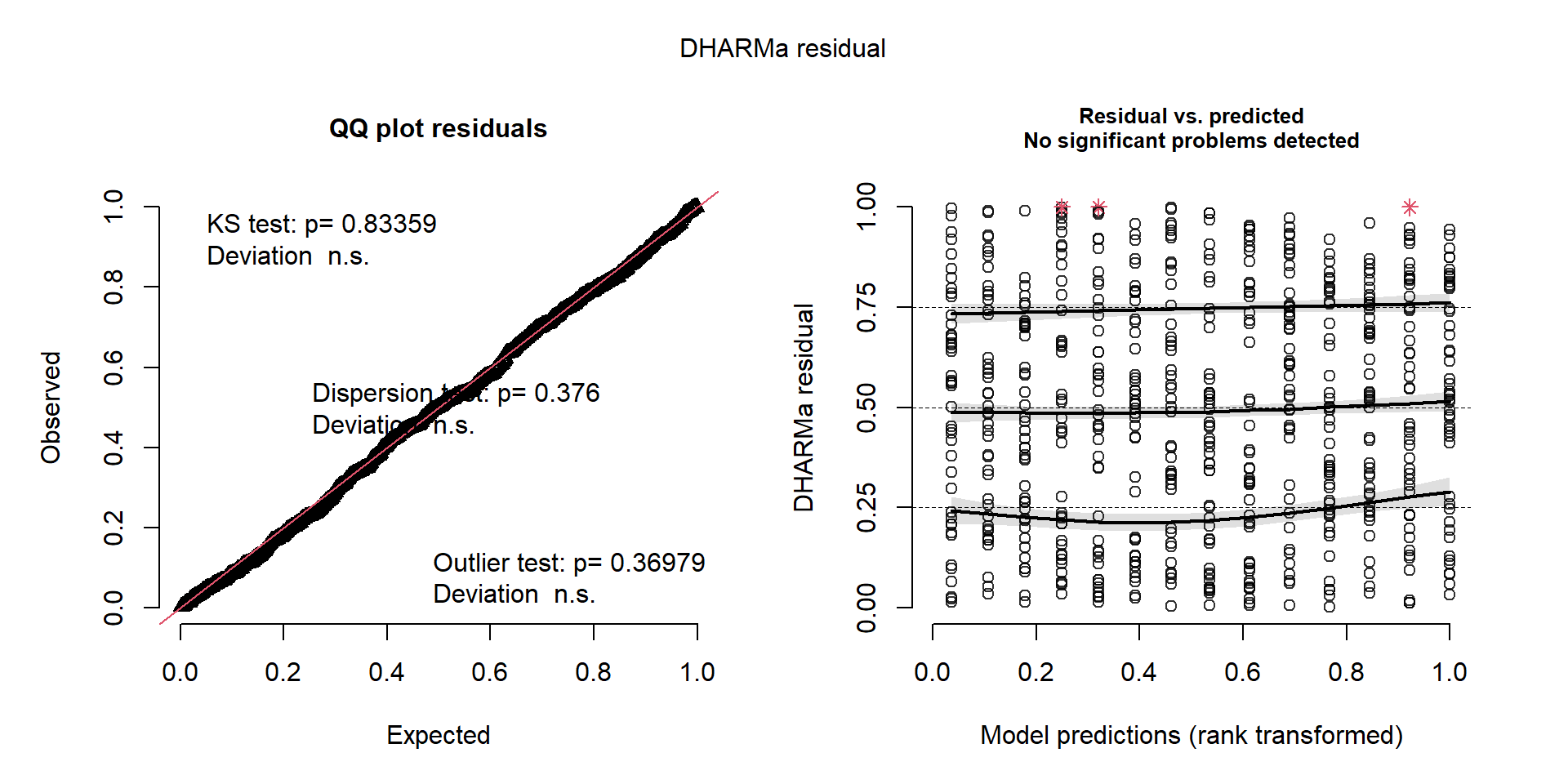
Excess zeros: ZI & hurdle models
- Zero-Inflated models:
- Some zeros are true, others are structural
- Zero-Inflated Poisson (ZIP): two processes (occurence)
- Zero-Inflated Negative Binomial (ZINB): overdispersion
- Hurdle models:
- Hurdle models are two-part models: 1) classification to detect zeros + 2) regression to model truncated data
- Here, all zeros are structural
Zero-Inflated Poisson model
m3_zip <- glmmTMB(count ~ spp + mined + (1|site), ziformula = ~mined + DOP + Wtemp,
data = Salamanders, family = poisson)
summary(m3_zip) Family: poisson ( log )
Formula: count ~ spp + mined + (1 | site)
Zero inflation: ~mined + DOP + Wtemp
Data: Salamanders
AIC BIC logLik deviance df.resid
1798.3 1856.4 -886.2 1772.3 631
Random effects:
Conditional model:
Groups Name Variance Std.Dev.
site (Intercept) 0.1281 0.3579
Number of obs: 644, groups: site, 23
Conditional model:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -0.4177 0.3009 -1.388 0.16512
sppPR -1.2680 0.2408 -5.266 1.39e-07 ***
sppDM 0.2690 0.1377 1.954 0.05074 .
sppEC-A -0.5619 0.2050 -2.741 0.00612 **
sppEC-L 0.6666 0.1278 5.216 1.83e-07 ***
sppDES-L 0.6229 0.1257 4.955 7.23e-07 ***
sppDF 0.1145 0.1471 0.778 0.43630
minedno 1.3286 0.2952 4.500 6.79e-06 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Zero-inflation model:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 0.75729 0.29674 2.552 0.0107 *
minedno -1.84122 0.33165 -5.552 2.83e-08 ***
DOP 0.13287 0.12488 1.064 0.2873
Wtemp -0.03289 0.12703 -0.259 0.7957
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Zero-Inflated: under the hood
zi_probs <- predict(m3_zip, type = "zprob") # Probability of excess zeros
ggplot(Salamanders, aes(x = mined, y = zi_probs)) +
geom_jitter(aes(color = as.factor(count == 0)), width = 0.5, size = 2, alpha = 0.6) +
stat_summary(fun = mean, geom = "point", color = "black", size = 5) +
theme_minimal() +
labs(x = "Mining Presence", y = "Probability of Excess Zeros", color = "Observed Zero?")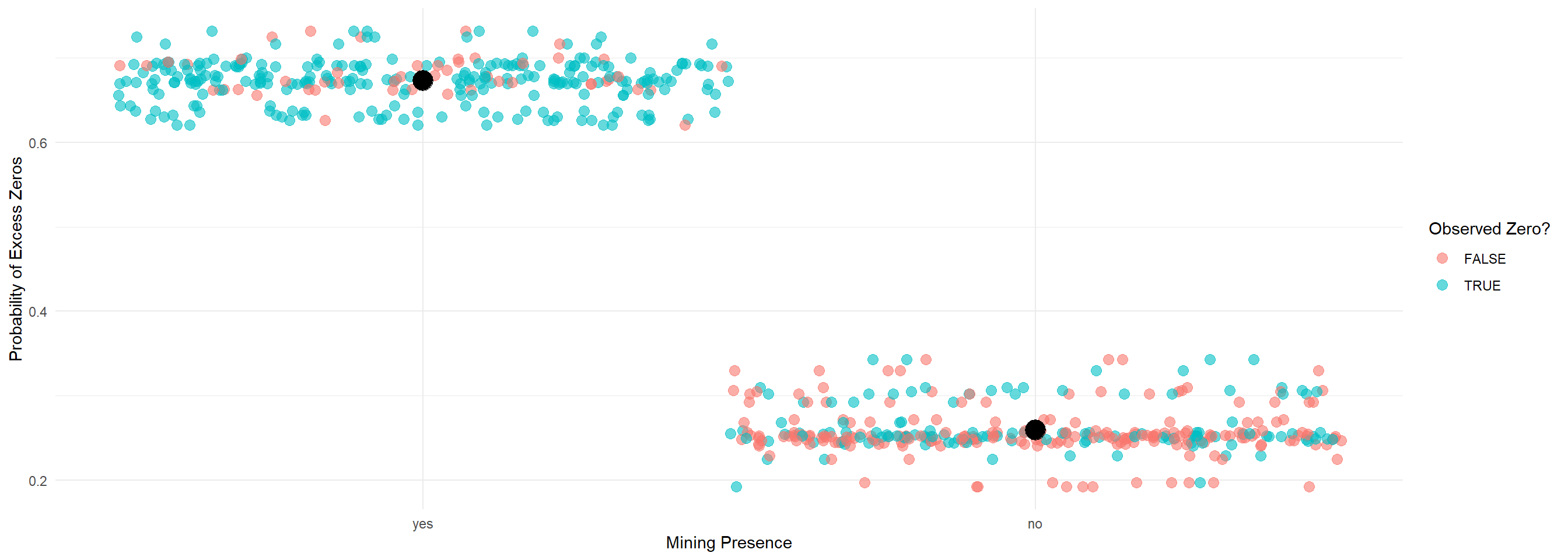
Zero-Inflated Negative Binomial model
m4_zinb <- glmmTMB(count ~ spp + mined + (1|site), ziformula = ~mined + DOP + Wtemp,
data = Salamanders, family = nbinom2)
summary(m4_zinb) Family: nbinom2 ( log )
Formula: count ~ spp + mined + (1 | site)
Zero inflation: ~mined + DOP + Wtemp
Data: Salamanders
AIC BIC logLik deviance df.resid
1668.0 1730.5 -820.0 1640.0 630
Random effects:
Conditional model:
Groups Name Variance Std.Dev.
site (Intercept) 0.1403 0.3745
Number of obs: 644, groups: site, 23
Dispersion parameter for nbinom2 family (): 1.05
Conditional model:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -0.8593 0.3839 -2.238 0.02522 *
sppPR -1.3781 0.2860 -4.818 1.45e-06 ***
sppDM 0.3084 0.2261 1.364 0.17260
sppEC-A -0.7569 0.2523 -3.000 0.00270 **
sppEC-L 0.5388 0.2203 2.445 0.01447 *
sppDES-L 0.7154 0.2194 3.261 0.00111 **
sppDF 0.1720 0.2355 0.730 0.46511
minedno 1.4999 0.3535 4.242 2.21e-05 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Zero-inflation model:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -0.3065 0.6593 -0.465 0.642
minedno -20.5209 3451.8717 -0.006 0.995
DOP -1.0964 0.7052 -1.555 0.120
Wtemp -0.2229 0.3414 -0.653 0.514Model comparison
- Compare models using AIC, residual plots, likelihood ratio tests
Data: Salamanders
Models:
m1_pr: count ~ spp + mined + (1 | site), zi=~0, disp=~1
m2_nb: count ~ spp + mined + (1 | site), zi=~0, disp=~1
m3_zip: count ~ spp + mined + (1 | site), zi=~mined + DOP + Wtemp, disp=~1
m4_zinb: count ~ spp + mined + (1 | site), zi=~mined + DOP + Wtemp, disp=~1
Df AIC BIC logLik deviance Chisq Chi Df Pr(>Chisq)
m1_pr 9 1962.8 2003.0 -972.40 1944.8
m2_nb 10 1672.4 1717.1 -826.20 1652.4 292.40 1 <2e-16 ***
m3_zip 13 1798.3 1856.4 -886.17 1772.3 0.00 3 1
m4_zinb 14 1668.0 1730.5 -820.00 1640.0 132.35 1 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Visualization code
# Generate predictions for all models
Salamanders <- Salamanders %>%
mutate(
pred_pr = predict(m1_pr, type = "response"),
pred_nb = predict(m2_nb, type = "response"),
pred_zip = predict(m3_zip, type = "response"),
pred_zinb = predict(m4_zinb, type = "response")
)
# Plot observed vs. predicted
ggplot(Salamanders, aes(x = count)) +
geom_histogram(binwidth = 1, alpha = 0.5, color = "white") +
geom_density(aes(x = pred_pr, y = ..count..), color = "#2980b9", size = 1.2, linetype = "dashed") +
geom_density(aes(x = pred_nb, y = ..count..), color = "#c0392b", size = 1.2, linetype = "dashed") +
geom_density(aes(x = pred_zip, y = ..count..), color = "#8e44ad", size = 1.2, linetype = "dotted") +
geom_density(aes(x = pred_zinb, y = ..count..), color = "#16a085", size = 1.2) +
theme_minimal() +
labs(
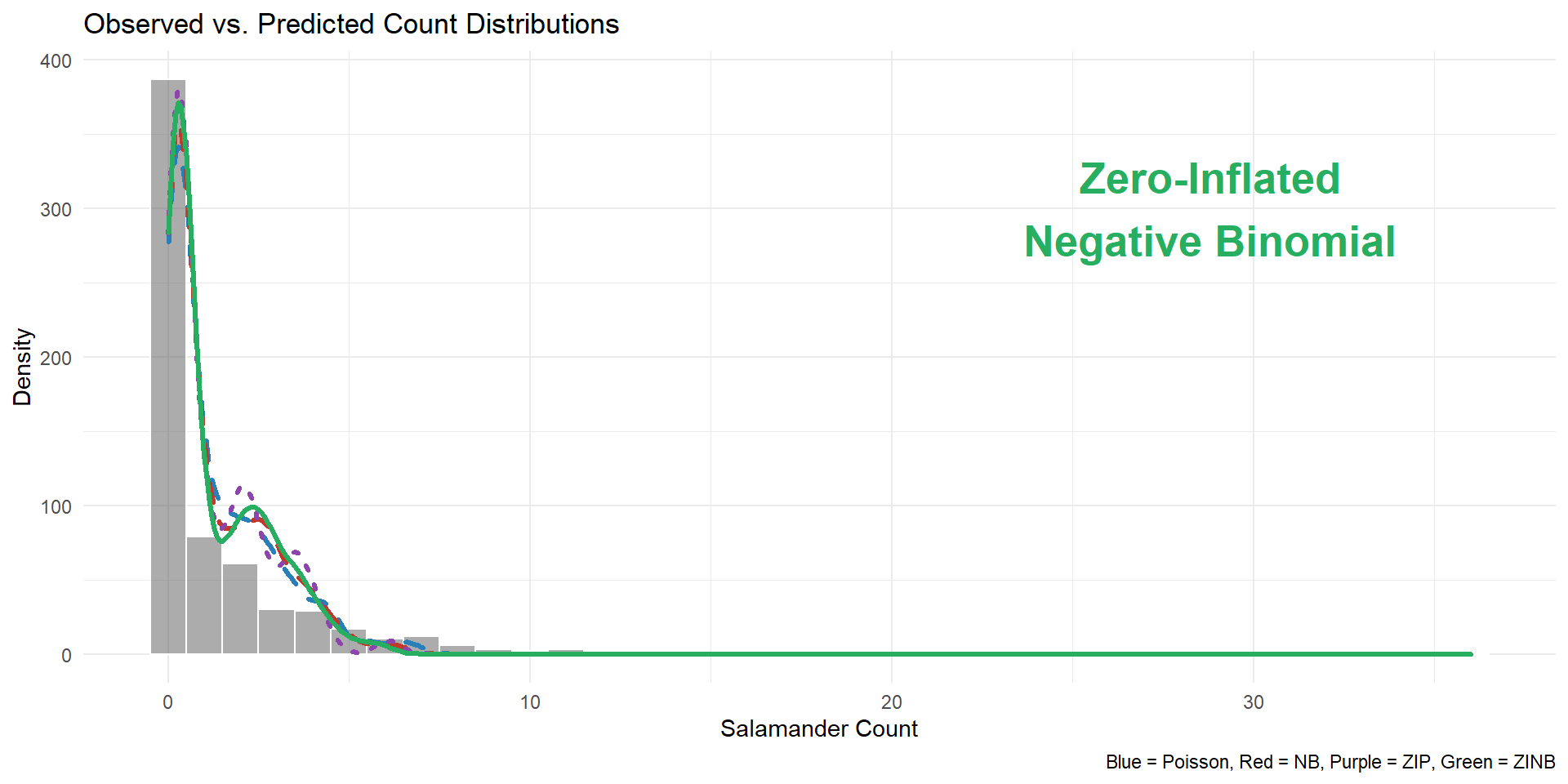
title = "Observed vs. Predicted Count Distributions",
x = "Salamander Count",
y = "Density",
caption = "Blue = Poisson, Red = NB, Purple = ZIP, Green = ZINB"
)Visualization output
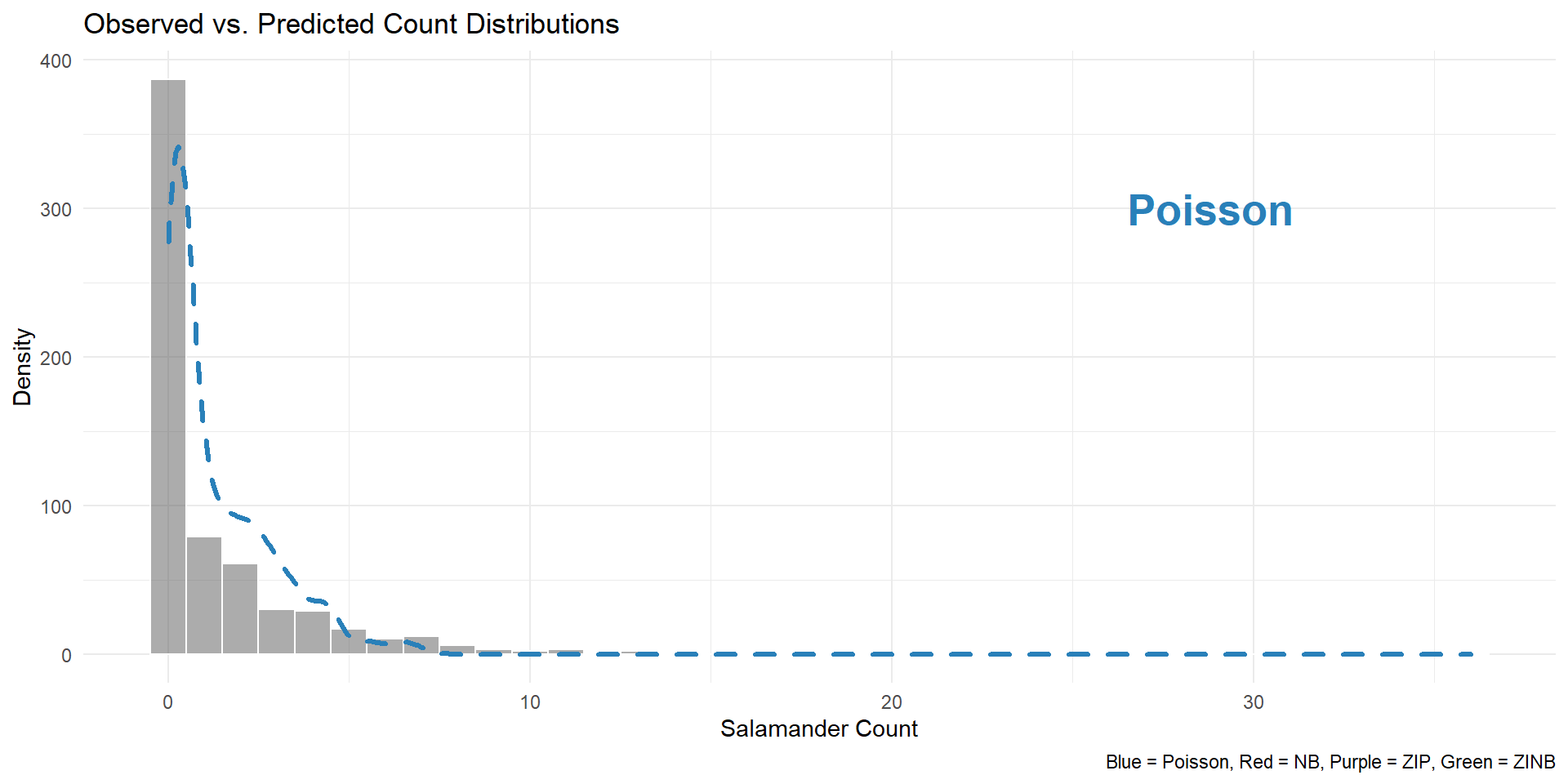
Visualization output
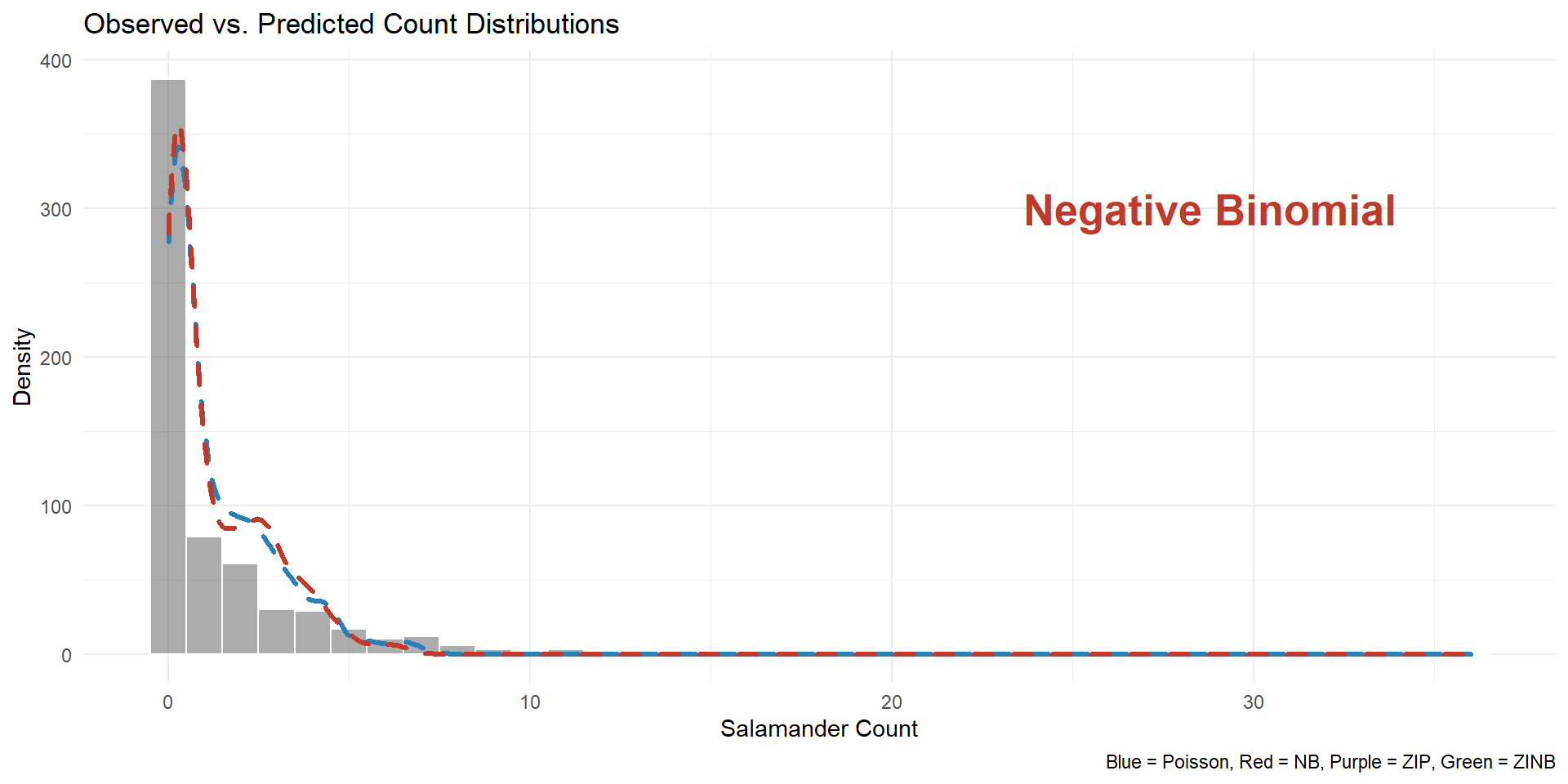
Visualization output
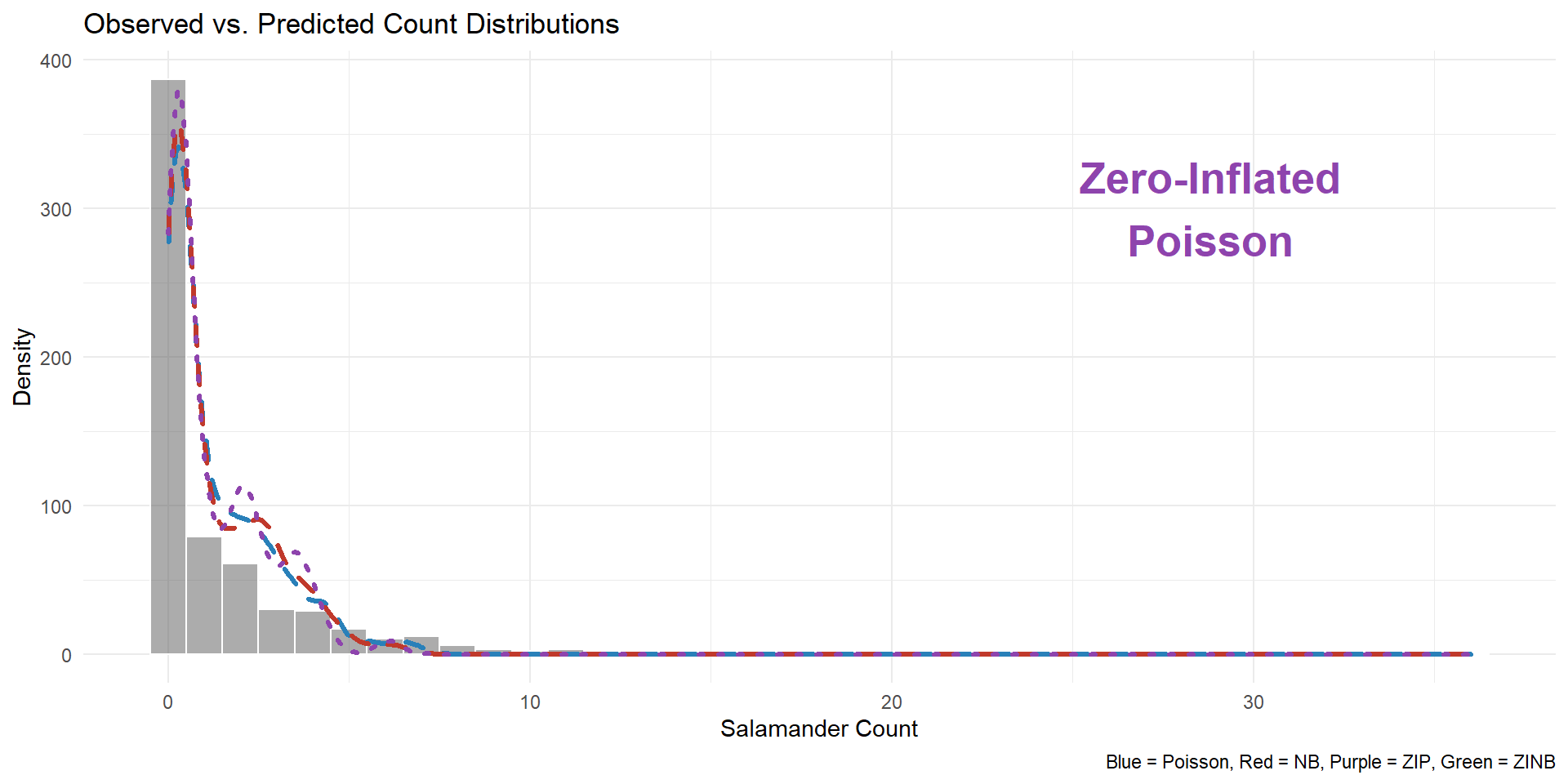
Visualization output
Thank you! Obrigado!
- github.com/Delvis/ZIPcode


